Description
| Product Name: | Rat PCNA (ProlifeRating Cell Nuclear Antigen) ELISA Kit |
| Product Code: | RTFI01032 |
| Size: | 96 Assays |
| Target: | Rat PCNA |
| Alias: | PCNA |
| Reactivity: | Rat |
| Detection Method: | Sandwich ELISA, Double Antibody |
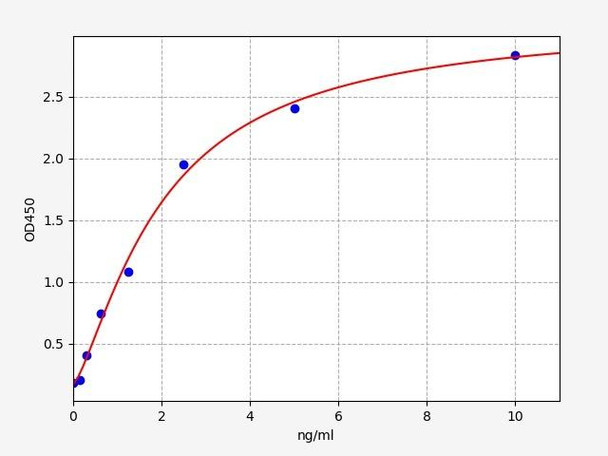
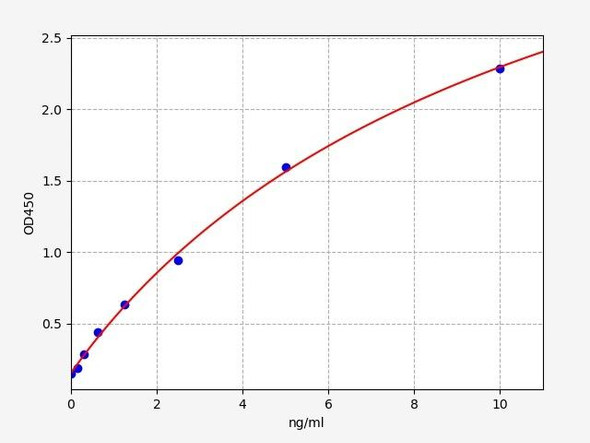
| Sensitivity: | 0.094ng/ml |
| Range: | 0.156-10ng/ml |
| Storage: | 4°C for 6 months |
| Note: | For Research Use Only |
| Recovery: | Matrices listed below were spiked with certain level of Rat PCNA and the recovery rates were calculated by comparing the measured value to the expected amount of Rat PCNA in samples. | ||||||||||||||||
| |||||||||||||||||
| Linearity: | The linearity of the kit was assayed by testing samples spiked with appropriate concentration of Rat PCNA and their serial dilutions. The results were demonstrated by the percentage of calculated concentration to the expected. | ||||||||||||||||
| |||||||||||||||||
| Intra-Assay: | CV <8% | ||||||||||||||||
| Inter-Assay: | CV <10% |
| Uniprot: | P04961 |
| UniProt Protein Function: | PCNA: This protein is an auxiliary protein of DNA polymerase delta and is involved in the control of eukaryotic DNA replication by increasing the polymerase's processibility during elongation of the leading strand. Induces a robust stimulatory effect on the 3'- 5' exonuclease and 3'-phosphodiesterase, but not apurinic- apyrimidinic (AP) endonuclease, APEX2 activities. Has to be loaded onto DNA in order to be able to stimulate APEX2. Homotrimer. Forms a complex with activator 1 heteropentamer in the presence of ATP. Interacts with EXO1, POLH, POLK, DNMT1, ERCC5, FEN1, CDC6 and POLDIP2. Interacts with APEX2; this interaction is triggered by reactive oxygen species and increased by misincorporation of uracil in nuclear DNA. Forms a ternary complex with DNTTIP2 and core histone. Interacts with KCTD10 and PPP1R15A. Interacts with POLD1, POLD3 and POLD4. Interacts with BAZ1B; the interaction is direct. Interacts with HLTF and SHPRH. Interacts with NUDT15. Interaction is disrupted in response to UV irradiation and acetylation. Interacts with p21Cip1/p21(CIP1) and CDT1; interacts via their PIP-box which also recruits the DCX(DTL) complex. Interacts with DDX11. Interacts with EGFR; positively regulates PCNA. Interacts with C12orf48/PARI. Interacts with SMARCAD1. Belongs to the PCNA family. |
| UniProt Protein Details: | Protein type:DNA replication; Cell cycle regulation Cellular Component: cell; centrosome; cyclin-dependent protein kinase holoenzyme complex; cytoplasm; intracellular; nuclear chromosome, telomeric region; nuclear lamina; nuclear replication fork; nucleoplasm; nucleus; PCNA complex; replication fork Molecular Function:chromatin binding; damaged DNA binding; dinucleotide insertion or deletion binding; DNA polymerase processivity factor activity; enzyme binding; estrogen receptor binding; histone acetyltransferase binding; identical protein binding; MutLalpha complex binding; protein binding; purine-specific mismatch base pair DNA N-glycosylase activity; receptor tyrosine kinase binding; transcription factor binding Biological Process: base-excision repair, gap-filling; bypass DNA synthesis; epithelial cell differentiation; heart development; leading strand elongation; mismatch repair; negative regulation of transcription from RNA polymerase II promoter; positive regulation of deoxyribonuclease activity; positive regulation of DNA repair; positive regulation of DNA replication; replication fork processing; response to cadmium ion; response to estradiol stimulus; response to lipid; response to oxidative stress |
| NCBI Summary: | cell cycle protein that may contribute to the repair of DNA damage and to cell proliferation caused by exposure to pulmonary toxicants; may influence cell survival in ischaemically compromised tissue [RGD, Feb 2006] |
| UniProt Code: | P04961 |
| NCBI GenInfo Identifier: | 129698 |
| NCBI Gene ID: | 25737 |
| NCBI Accession: | P04961.1 |
| UniProt Related Accession: | P04961 |
| Molecular Weight: | 28,749 Da |
| NCBI Full Name: | Proliferating cell nuclear antigen |
| NCBI Synonym Full Names: | proliferating cell nuclear antigen |
| NCBI Official Symbol: | Pcna |
| NCBI Official Synonym Symbols: | PCNAR; Pcna/cyclin |
| NCBI Protein Information: | proliferating cell nuclear antigen |
| UniProt Protein Name: | Proliferating cell nuclear antigen |
| UniProt Synonym Protein Names: | Cyclin |
| Protein Family: | PCNA-associated factor |
| UniProt Gene Name: | Pcna |
| UniProt Entry Name: | PCNA_RAT |
| Step | Procedure |
| 1. | Set standard, test sample and control (zero) wells on the pre-coated plate respectively, and then, record their positions. It is recommended to measure each standard and sample in duplicate. Wash plate 2 times before adding standard, sample and control (zero) wells! |
| 2. | Aliquot 0.1ml standard solutions into the standard wells. |
| 3. | Add 0.1 ml of Sample / Standard dilution buffer into the control (zero) well. |
| 4. | Add 0.1 ml of properly diluted sample ( Human serum, plasma, tissue homogenates and other biological fluids.) into test sample wells. |
| 5. | Seal the plate with a cover and incubate at 37°C for 90 min. |
| 6. | Remove the cover and discard the plate content, clap the plate on the absorbent filter papers or other absorbent material. Do NOT let the wells completely dry at any time. Wash plate X2. |
| 7. | Add 0.1 ml of Biotin- detection antibody working solution into the above wells (standard, test sample & zero wells). Add the solution at the bottom of each well without touching the side wall. |
| 8. | Seal the plate with a cover and incubate at 37°C for 60 min. |
| 9. | Remove the cover, and wash plate 3 times with Wash buffer. Let wash buffer rest in wells for 1 min between each wash. |
| 10. | Add 0.1 ml of SABC working solution into each well, cover the plate and incubate at 37°C for 30 min. |
| 11. | Remove the cover and wash plate 5 times with Wash buffer, and each time let the wash buffer stay in the wells for 1-2 min. |
| 12. | Add 90 µL of TMB substrate into each well, cover the plate and incubate at 37°C in dark within 10-20 min. (Note: This incubation time is for reference use only, the optimal time should be determined by end user.) And the shades of blue can be seen in the first 3-4 wells (with most concentrated standard solutions), the other wells show no obvious color. |
| 13. | Add 50 µL of Stop solution into each well and mix thoroughly. The color changes into yellow immediately. |
| 14. | Read the O.D. absorbance at 450 nm in a microplate reader immediately after adding the stop solution. |
When carrying out an ELISA assay it is important to prepare your samples in order to achieve the best possible results. Below we have a list of procedures for the preparation of samples for different sample types.
| Sample Type | Protocol |
| Serum: | If using serum separator tubes, allow samples to clot for 30 minutes at room temperature. Centrifuge for 10 minutes at 1,000x g. Collect the serum fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. If serum separator tubes are not being used, allow samples to clotovernight at 2-8°C. Centrifuge for 10 minutes at 1,000x g. Removeserum and assay promptly or aliquot and store the samples at-80°C. Avoid multiple freeze-thaw cycles. |
| Plasma: | Collect plasma using EDTA or heparin as an anti-coagulant. Centrifuge samples at 4°C for 15 mins at 1000 × g within 30 mins of collection. Collect the plasma fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles.Note: Over haemolysed samples are not suitable for use with this kit. |
| Urine & Cerebrospinal Fluid: | Collect the urine (mid-stream) in a sterile container, centrifuge for 20 mins at 2000-3000 rpm. Remove supernatant and assay immediately. If any precipitation is detected, repeat the centrifugation step. A similar protocol can be used for cerebrospinal fluid. |
| Cell Culture Supernatant: | Collect the cell culture media by pipette, followed by centrifugation at 4°C for 20 mins at 1500 rpm. Collect the clear supernatant and assay immediately. |
| Cell Lysates: | Solubilize cells in lysis buffer and allow to sit on ice for 30 minutes. Centrifuge tubes at 14,000 x g for 5 minutes to remove insoluble material. Aliquot the supernatant into a new tube and discard the remaining whole cell extract. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20°C. |
| Tissue Homogenates: | The preparation of tissue homogenates will vary depending upon tissue type. Rinse tissue with 1X PBS to remove excess blood & homogenizein 20ml of 1X PBS (including protease inhibitors) and store overnight at ≤ -20°C. Two freeze-thaw cycles are required to break the cell membranes. To further disrupt the cell membranes you can sonicate the samples. Centrifuge homogenates for 5 mins at 5000xg. Remove the supernatant and assay immediately or aliquot and store at -20°C or-80°C. |
| Tissue Lysates: | Rinse tissue with PBS, cut into 1-2 mm pieces, and homogenize with a tissue homogenizer in PBS. Add an equal volume of RIPA buffer containing protease inhibitors and lyse tissues at room temperature for 30 minutes with gentle agitation. Centrifuge to remove debris. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Breast Milk: | Collect milk samples and centrifuge at 10,000 x g for 60 min at 4°C. Aliquot the supernatant and assay. For long term use, store samples at -80°C. Minimize freeze/thaw cycles. |
Additional Information
Product Type: |
ELISA Kit |
Size: |
96 Assays |
Uniprot: |
P04961 |
Sensitivity: |
0.094ng/ml |
Range: |
0.156-10ng/ml |
ELISA Type: |
Sandwich |
Synonyms: |
PCNA |
Reactivity: |
Rat |
Research Area: |
Epigenetics and Nuclear Signaling |