Description
| Product Name: | Human SMN1 / Survival of Motor Neuron 1, Telomeric ELISA Kit |
| Product Code: | HUFI02863 |
| Size: | 96 Assays |
| Alias: | SMN1, Survival of Motor Neuron 1, Telomeric |
| Detection method: | Sandwich ELISA, Double Antibody |
| Application: | This immunoassay kit allows for the in vitro quantitative determination of Human SMN1 concentrations in serum plasma and other biological fluids. |
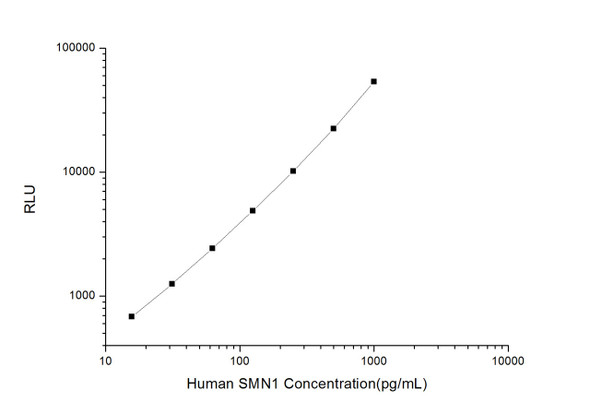
| Sensitivity: | 18.75pg/ml |
| Range: | 31.25-2000pg/ml |
| Storage: | 4°C for 6 months |
| Note: | For Research Use Only |
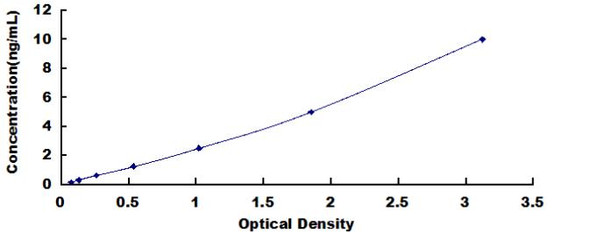
| Recovery: | Matrices listed below were spiked with certain level of Human SMN1 and the recovery rates were calculated by comparing the measured value to the expected amount of Human SMN1 in samples. | ||||||||||||||||
| |||||||||||||||||
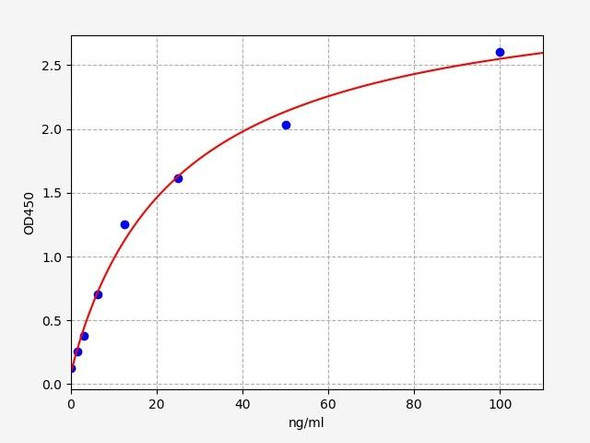
| Linearity: | The linearity of the kit was assayed by testing samples spiked with appropriate concentration of Human SMN1 and their serial dilutions. The results were demonstrated by the percentage of calculated concentration to the expected. | ||||||||||||||||
| |||||||||||||||||
| CV(%): | Intra-Assay: CV<8% Inter-Assay: CV<10% |
| Component | Quantity | Storage |
| ELISA Microplate (Dismountable) | 8×12 strips | 4°C for 6 months |
| Lyophilized Standard | 2 | 4°C/-20°C |
| Sample/Standard Dilution Buffer | 20ml | 4°C |
| Biotin-labeled Antibody(Concentrated) | 120ul | 4°C (Protect from light) |
| Antibody Dilution Buffer | 10ml | 4°C |
| HRP-Streptavidin Conjugate(SABC) | 120ul | 4°C (Protect from light) |
| SABC Dilution Buffer | 10ml | 4°C |
| TMB Substrate | 10ml | 4°C (Protect from light) |
| Stop Solution | 10ml | 4°C |
| Wash Buffer(25X) | 30ml | 4°C |
| Plate Sealer | 5 | - |
Other materials and equipment required:
- Microplate reader with 450 nm wavelength filter
- Multichannel Pipette, Pipette, microcentrifuge tubes and disposable pipette tips
- Incubator
- Deionized or distilled water
- Absorbent paper
- Buffer resevoir
| Uniprot | Q16637 |
| UniProt Protein Function: | SMN: The SMN complex plays an essential role in spliceosomal snRNP assembly in the cytoplasm and is required for pre-mRNA splicing in the nucleus. It may also play a role in the metabolism of snoRNPs. Defects in SMN1 are the cause of spinal muscular atrophy autosomal recessive type 1 (SMA1). Spinal muscular atrophy refers to a group of neuromuscular disorders characterized by degeneration of the anterior horn cells of the spinal cord, leading to symmetrical muscle weakness and atrophy. Autosomal recessive forms are classified according to the age of onset, the maximum muscular activity achieved, and survivorship. The severity of the disease is mainly determined by the copy number of SMN2, a copy gene which predominantly produces exon 7-skipped transcripts and only low amount of full-length transcripts that encode for a protein identical to SMN1. Only about 4% of SMA patients bear one SMN1 copy with an intragenic mutation. SMA1 is a severe form, with onset before 6 months of age. SMA1 patients never achieve the ability to sit. Defects in SMN1 are the cause of spinal muscular atrophy autosomal recessive type 2 (SMA2). SMA2 is an autosomal recessive spinal muscular atrophy of intermediate severity, with onset between 6 and 18 months. Patients do not reach the motor milestone of standing, and survive into adulthood. Defects in SMN1 are the cause of spinal muscular atrophy autosomal recessive type 3 (SMA3). SMA3 is an autosomal recessive spinal muscular atrophy with onset after 18 months. SMA3 patients develop ability to stand and walk and survive into adulthood. Defects in SMN1 are the cause of spinal muscular atrophy autosomal recessive type 4 (SMA4). SMA4 is an autosomal recessive spinal muscular atrophy characterized by symmetric proximal muscle weakness with onset in adulthood and slow disease progression. SMA4 patients can stand and walk. Belongs to the SMN family. 4 isoforms of the human protein are produced by alternative splicing. |
| UniProt Protein Details: | Protein type:RNA processing; RNA-binding Chromosomal Location of Human Ortholog: 5q13.2 Cellular Component: Cajal body; cytoplasm; cytosol; neuron projection; nucleoplasm; nucleus; perikaryon; SMN complex Molecular Function:identical protein binding; protein binding Biological Process: nuclear import; spliceosomal snRNP biogenesis; spliceosome assembly; transcription termination Disease: Spinal Muscular Atrophy, Type I; Spinal Muscular Atrophy, Type Ii; Spinal Muscular Atrophy, Type Iii; Spinal Muscular Atrophy, Type Iv |
| NCBI Summary: | This gene is part of a 500 kb inverted duplication on chromosome 5q13. This duplicated region contains at least four genes and repetitive elements which make it prone to rearrangements and deletions. The repetitiveness and complexity of the sequence have also caused difficulty in determining the organization of this genomic region. The telomeric and centromeric copies of this gene are nearly identical and encode the same protein. While mutations in the telomeric copy are associated with spinal muscular atrophy, mutations in this gene, the centromeric copy, do not lead to disease. This gene may be a modifier of disease caused by mutation in the telomeric copy. The critical sequence difference between the two genes is a single nucleotide in exon 7, which is thought to be an exon splice enhancer. Note that the nine exons of both the telomeric and centromeric copies are designated historically as exon 1, 2a, 2b, and 3-8. It is thought that gene conversion events may involve the two genes, leading to varying copy numbers of each gene. The full length protein encoded by this gene localizes to both the cytoplasm and the nucleus. Within the nucleus, the protein localizes to subnuclear bodies called gems which are found near coiled bodies containing high concentrations of small ribonucleoproteins (snRNPs). This protein forms heteromeric complexes with proteins such as SIP1 and GEMIN4, and also interacts with several proteins known to be involved in the biogenesis of snRNPs, such as hnRNP U protein and the small nucleolar RNA binding protein. Four transcript variants encoding distinct isoforms have been described. [provided by RefSeq, Sep 2008] |
| UniProt Code: | Q16637 |
| NCBI GenInfo Identifier: | 2498924 |
| NCBI Gene ID: | 6607 |
| NCBI Accession: | Q16637.1 |
| UniProt Secondary Accession: | Q16637,Q13119, Q549U5, Q96J51, A8K0V4, |
| UniProt Related Accession: | Q16637 |
| Molecular Weight: | 32kDa |
| NCBI Full Name: | Survival motor neuron protein |
| NCBI Synonym Full Names: | survival of motor neuron 2, centromeric |
| NCBI Official Symbol: | SMN2 |
| NCBI Official Synonym Symbols: | SMNC; BCD541; GEMIN1; TDRD16B; C-BCD541 |
| NCBI Protein Information: | survival motor neuron protein |
| UniProt Protein Name: | Survival motor neuron protein |
| UniProt Synonym Protein Names: | Component of gems 1; Gemin-1 |
| UniProt Gene Name: | SMN1 |
*Note: Protocols are specific to each batch/lot. For the correct instructions please follow the protocol included in your kit.
Before adding to wells, equilibrate the SABC working solution and TMB substrate for at least 30 min at 37°C. When diluting samples and reagents, they must be mixed completely and evenly. It is recommended to plot a standard curve for each test.
| Step | Protocol |
| 1. | Set standard, test sample and control (zero) wells on the pre-coated plate respectively, and then, record their positions. It is recommended to measure each standard and sample in duplicate. Wash plate 2 times before adding standard, sample and control (zero) wells! |
| 2. | Aliquot 0.1ml standard solutions into the standard wells. |
| 3. | Add 0.1 ml of Sample / Standard dilution buffer into the control (zero) well. |
| 4. | Add 0.1 ml of properly diluted sample ( Human serum, plasma, tissue homogenates and other biological fluids.) into test sample wells. |
| 5. | Seal the plate with a cover and incubate at 37 °C for 90 min. |
| 6. | Remove the cover and discard the plate content, clap the plate on the absorbent filter papers or other absorbent material. Do NOT let the wells completely dry at any time. Wash plate X2. |
| 7. | Add 0.1 ml of Biotin- detection antibody working solution into the above wells (standard, test sample & zero wells). Add the solution at the bottom of each well without touching the side wall. |
| 8. | Seal the plate with a cover and incubate at 37°C for 60 min. |
| 9. | Remove the cover, and wash plate 3 times with Wash buffer. Let wash buffer rest in wells for 1 min between each wash. |
| 10. | Add 0.1 ml of SABC working solution into each well, cover the plate and incubate at 37°C for 30 min. |
| 11. | Remove the cover and wash plate 5 times with Wash buffer, and each time let the wash buffer stay in the wells for 1-2 min. |
| 12. | Add 90 µl of TMB substrate into each well, cover the plate and incubate at 37°C in dark within 10-20 min. (Note: This incubation time is for reference use only, the optimal time should be determined by end user.) And the shades of blue can be seen in the first 3-4 wells (with most concentrated standard solutions), the other wells show no obvious color. |
| 13. | Add 50 µl of Stop solution into each well and mix thoroughly. The color changes into yellow immediately. |
| 14. | Read the O.D. absorbance at 450 nm in a microplate reader immediately after adding the stop solution. |
When carrying out an ELISA assay it is important to prepare your samples in order to achieve the best possible results. Below we have a list of procedures for the preparation of samples for different sample types.
| Sample Type | Protocol |
| Serum | If using serum separator tubes, allow samples to clot for 30 minutes at room temperature. Centrifuge for 10 minutes at 1,000x g. Collect the serum fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. If serum separator tubes are not being used, allow samples to clot overnight at 2-8°C. Centrifuge for 10 minutes at 1,000x g. Remove serum and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. |
| Plasma | Collect plasma using EDTA or heparin as an anticoagulant. Centrifuge samples at 4°C for 15 mins at 1000 × g within 30 mins of collection. Collect the plasma fraction and assay promptly or aliquot and store the samples at -80°C. Avoid multiple freeze-thaw cycles. Note: Over haemolysed samples are not suitable for use with this kit. |
| Urine & Cerebrospinal Fluid | Collect the urine (mid-stream) in a sterile container, centrifuge for 20 mins at 2000-3000 rpm. Remove supernatant and assay immediately. If any precipitation is detected, repeat the centrifugation step. A similar protocol can be used for cerebrospinal fluid. |
| Cell culture supernatant | Collect the cell culture media by pipette, followed by centrifugation at 4°C for 20 mins at 1500 rpm. Collect the clear supernatant and assay immediately. |
| Cell lysates | Solubilize cells in lysis buffer and allow to sit on ice for 30 minutes. Centrifuge tubes at 14,000 x g for 5 minutes to remove insoluble material. Aliquot the supernatant into a new tube and discard the remaining whole cell extract. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Tissue homogenates | The preparation of tissue homogenates will vary depending upon tissue type. Rinse tissue with 1X PBS to remove excess blood & homogenize in 20ml of 1X PBS (including protease inhibitors) and store overnight at ≤ -20°C. Two freeze-thaw cycles are required to break the cell membranes. To further disrupt the cell membranes you can sonicate the samples. Centrifuge homogenates for 5 mins at 5000xg. Remove the supernatant and assay immediately or aliquot and store at -20°C or -80°C. |
| Tissue lysates | Rinse tissue with PBS, cut into 1-2 mm pieces, and homogenize with a tissue homogenizer in PBS. Add an equal volume of RIPA buffer containing protease inhibitors and lyse tissues at room temperature for 30 minutes with gentle agitation. Centrifuge to remove debris. Quantify total protein concentration using a total protein assay. Assay immediately or aliquot and store at ≤ -20 °C. |
| Breast Milk | Collect milk samples and centrifuge at 10,000 x g for 60 min at 4°C. Aliquot the supernatant and assay. For long term use, store samples at -80°C. Minimize freeze/thaw cycles. |
Fill out our quote form below and a dedicated member of staff will get back to you within one working day!
Additional Information
Product Type: |
ELISA Kit |
Size: |
96 Assays |
Uniprot: |
Q16637 |
Sensitivity: |
18.75pg/ml |
Range: |
31.25-2000pg/ml |
ELISA Type: |
Sandwich |
Synonyms: |
SMN1, Survival of Motor Neuron 1, Telomeric |
Reactivity: |
Human |